Most of our research is into how the brain balances its energy supply with demand – and how variations in this balance affect brain function.
Specifically, we want to learn which vascular cells control the brain’s energy supply, and how much neurovascular coupling between energy supply (blood flow) and demand (neuronal activity) varies physiologically and during disease states. For example, we are investigating whether neurovascular function changes during conditions such as Alzheimer’s disease, COVID-19 and obesity, across brain states and brain regions. To do this, we image blood vessels in brain slices and in vivo, in behaving mice, while manipulating neuronal activity pharmacologically or using physiological stimuli, and imaging neuronal activity using genetically encoded calcium indicators. Our results will be important for understanding how brain activity is fuelled and how the brain’s energy supply is affected by disease. Because human functional magnetic resonance imaging (fMRI) measures blood flow as a surrogate of neuronal activity, our results will also be important for understanding what fMRI signals actually tell us about neuronal activity.
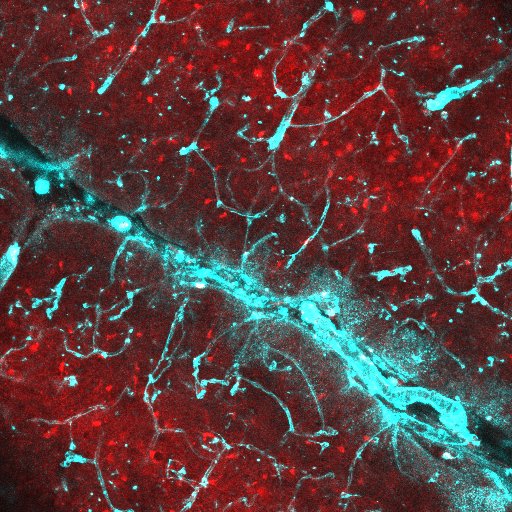
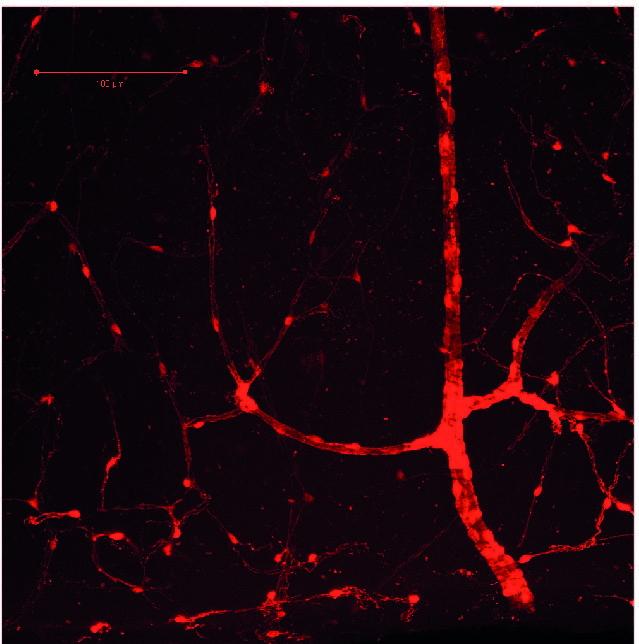
Brain blood vessels can be imaged in behaving mice, by intravenously injecting a fluorescent dye and imaging the brain with a 2-photon microscope (red in movie above). We can also see what the neurons are doing, by labelling them with a fluorescent calcium indicator (GCaMP6f; green). As the mouse runs on a wheel, and watches visual stimuli, neurons are activated and blood vessels dilate. You can see by eye the large vessel in the centre dilate, but when we take higher resolution movies of the smaller capillaries, we can also see them dilate.
We can also study neurovascular function in live or fixed brain slices.
Projects
How does neurovascular coupling differ across brain regions?
The brain needs energy to fuel its activity. Active neurons produce “neurovascular coupling” signals, which diffuse to blood vessels to increase blood flow, and supply oxygen and glucose. The undersupply of oxygen and glucose to the brain is associated with neuronal damage (e.g. Alzheimer’s disease). Neurovascular coupling has been widely studied in primary sensory cortices in vivo, but less so in hippocampus. Evidence has suggested that neuronal activity can occur without a positive BOLD signal in hippocampus 1,2, and that this region is uniquely more vulnerable to hypoxia 3. Why and when such “neurovascular uncoupling” occurs in the hippocampus is unclear.
We used two-photon imaging to visualise blood vessels and excitatory neurons (A) in the dorsal hippocampus in vivo from awake mice traversing a virtual reality environment (B). Image A shows the pyramidal cell layer (stratum pyramidale) of CA1, with active excitatory neurons labelled in green (GCaMP6f) and blood vessels in fluorescent red (Texas red dextran). We found lower blood flow, blood oxygenation and weaker neurovascular coupling in the hippocampus than in the cortex 4. We are now studying neurovascular function in the corpus callosum using similar techniques, as well as trying to understand how hippocampus function may be affected by these differences during the emergence of disease.

1 – Mishra et al. J. Neurosci. 31, 15053-64, (2011). 2 – Angenstein et al. J. Neurosci. 29, 2428-39, (2009). 3 - Cervos-Navarro & Diemer. Crit Re Neurobiol. 6, 149-82, (1991). 4 - Shaw et al. Nature Communications. 12, 3190, (2021).
How do Alzheimer’s disease risk factors affect neurovascular function? And how does neurovascular dysfunction affect neuronal and glial function?
We are interested in whether disrupted blood flow that occurs in advance of Alzheimer’s disease is important for driving the emergence of disease. To try and understand this, we have asked whether the main genetic risk factor for Alzheimer’s disease, the apolipoprotein e4 (APOE4) gene, disrupts neurovascular regulation. We find that on its own APOE4 only subtly neurovascular function 5 , but additional stressors such as physical inactivity significantly increase dysfunction. We are now investigating how beta amyloid interacts with these effects. Conversely we are also trying to understand how alterations in the brain’s blood supply affects the activity of different cell types, to understand how disrupted blood flow could promote disease.
5 – Bonnar et al. JCBFM 43(11):1826-1841 (2023)

Neuronal coding in the hippocampus during health and disease
The hippocampus is a brain area that is important for spatial navigation and spatial memory. It contains excitatory pyramidal cells that fire at a specific place in the environment, aptly named ‘place cells’. Although place cells are relatively stable within the same environment, they remap when part of the environment is changed. This might be a chance in local cues, e.g. objects added or removed, or to the context, e.g. a different scent is introduced. Not all place cells remap in the same way, some cells might start firing in a different location, others might stop firing alltogether. How place cells are defined in a population has a big effect on what properties they appear to have 6.
By combining two-photon imaging and presentation of a virtual environment, we study the individual response of cells to small changes to the environment. We use this to determine how individual aspects of the environment are encoded and study the functional heterogeneity of the place cells population, as well as how this changes during disease.
